Researchers at the University of Maryland use an all-atom lipid force field to create the first simulations of the soybean plasma membrane.
From the Journal: The Journal of Chemical Physics
WASHINGTON, D.C., June 6, 2017 — A cell’s plasma membrane forms a protective barrier, separating its inner contents from the outside environment. There is a pressing need to better understand the complex lipid bilayer that makes up this membrane, which limits the molecules that can leave or enter the cell. Research into the plasma membrane structure and behavior can provide invaluable information about whether, and to what extent, small molecules such as sugars, hormones and drugs, can permeate.
Researchers at the University of Maryland, in College Park, have developed a detailed computational model of the soybean plasma membrane that provides new structural insight at the molecular level. The results of their large-scale simulations highlight unique properties of the soybean plasma membrane and demonstrate a microscale membrane structure in which similar lipids tend to cluster together.
This novel research has applications for studying membrane proteins, which may be useful for engineering plants to produce biochemicals, biofuels, drugs and other compounds, and in understanding how plants sense and respond to stressful conditions. The group published their findings this week in The Journal of Chemical Physics, from AIP Publishing.
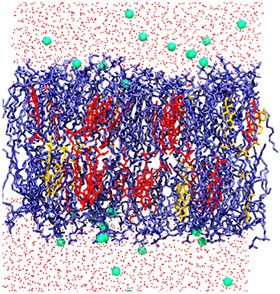 Most research on modeling plasma membranes has focused on single-celled microbes, such as E. coli or yeast, or on certain organs in model mammalian species. Both the bacteria and higher-level organisms have a double layer cell membrane composed of phospholipids, where the hydrophobic tails of each layer point toward the middle of the membrane and hydrophilic heads face the outside and inside of the cell. Depending on their concentration, sterol molecules can enhance membrane fluidity or increase its rigidity.
Most research on modeling plasma membranes has focused on single-celled microbes, such as E. coli or yeast, or on certain organs in model mammalian species. Both the bacteria and higher-level organisms have a double layer cell membrane composed of phospholipids, where the hydrophobic tails of each layer point toward the middle of the membrane and hydrophilic heads face the outside and inside of the cell. Depending on their concentration, sterol molecules can enhance membrane fluidity or increase its rigidity.
The researchers focused on the soybean plasma membrane because it is one of the most intensely studied plant membranes, which provided substantial experimental data to be used in validating the computational model.
“Plant plasma membranes haven’t been studied before at the all-atom computation level,” said Jeffery Klauda, associate professor of chemical and biomolecular engineering at the University of Maryland and lead investigator of the work. “These plasma membranes contain proteins that are involved in controlling what goes in and out of the cell, so to look at those proteins that reside in the membrane, we need to understand what the membrane is.”
The researchers used molecular dynamics computer simulations to simulate the structure and dynamics of the complex lipid membrane, which used Newton’s equations of motion to understand how molecules move in response to forces generated by atomic interactions. Specifically, they used the all-atom CHARRM36 lipid force field to predict how lipids self-assemble into a bilayer membrane, using seven or eight of the main phospholipid types and two primary sterols found in soybean seedling membranes.
The model demonstrated good agreement with experimental measurements of the membrane and revealed physical differences between soybean membrane and previous models of membranes found in yeast and E. coli. The soybean membrane is about as rigid as the yeast membrane, but twice as rigid as the sterol-deficient E. coli cytoplasmic membrane.
The soybean model also showed that lipids with similar amounts of unsaturation tended to cluster together, behavior scientists had not previously observed for these plant lipids. The surprising clustering behavior was attributed to van der Waals interactions between the hydrophobic tails of the phospholipids.
In future work, Klauda and his colleagues plan to examine membranes from other plants. They also plan to model transport proteins that span the lipid bilayer and other proteins critical to membrane function. While these simulations represent the state-of-the-art in computational modeling of complex lipid membranes, Klauda acknowledges that he would have liked to include a greater diversity of lipid types in the simulation, as plant membranes can be composed of hundreds of different lipids, but the model could only accommodate the 10 most dominant ones.
“We are in a ripening field in which we have the ability to simulate and probe biologically relevant membranes,” Klauda concluded. “If we compare what we’ve done to what was done five to ten years ago, when membranes were represented by one or two lipids, we clearly see here that if you want to understand the structure of the membrane, you really need to include the diversity that exists in biology.”
###
For More Information:
Julia Majors
media@aip.org
301-209-3090
@AIPPhysicsNews
Article Title
Authors
Xiaohong Zhuang, Anna Ou and Jeffery Klauda
Author Affiliations
University of Maryland
