Template-assisted replication, which helps polymers grow longer while passing on sequences from generation to generation, could have enabled jump from monomers to self-replicating polymers
From the Journal: The Journal of Chemical Physics
WASHINGTON, D.C., July 28, 2015 — When life on Earth began nearly 4 billion years ago, long before humans, dinosaurs or even the earliest single-celled forms of life roamed, it may have started as a hiccup rather than a roar: small, simple molecular building blocks known as “monomers” coming together into longer “polymer” chains and falling apart in the warm pools of primordial ooze over and over again.
Then, somewhere along the line, these growing polymer chains developed the ability to make copies of themselves. Competition between these molecules would allow the ones most efficient at making copies of themselves to do so faster or with greater abundance, a trait that would be shared by the copies they made. These rapid replicators would fill the soup faster than the other polymers, allowing the information they encoded to be passed on from one generation to another and, eventually, giving rise to what we think of today as life.
Or so the story goes. But with no fossil record to check from those early days, it’s a narrative that still has some chapters missing. One question in particular remains problematic: what enabled the leap from a primordial soup of individual monomers to self-replicating polymer chains?
A new model published this week in The Journal of Chemical Physics, from AIP Publishing, proposes a potential mechanism by which self-replication could have emerged. It posits that template-assisted ligation, the joining of two polymers by using a third, longer one as a template, could have enabled polymers to become self-replicating.
“We tried to fill this gap in understanding between simple physical systems to something that can behave in a life-like manner and transmit information,” said Alexei Tkachenko, a researcher at Brookhaven National Laboratory. Tkachenko carried out the research alongside Sergei Maslov, a professor at the University of Illinois at Urbana-Champaign with joint appointment at Brookhaven.
Origins of Self-Replication
Self-replication is a complicated process — DNA, the basis for life on earth today, requires a coordinated cohort of enzymes and other molecules in order to duplicate itself. Early self-replicating systems were surely more rudimentary, but their existence in the first place is still somewhat baffling.
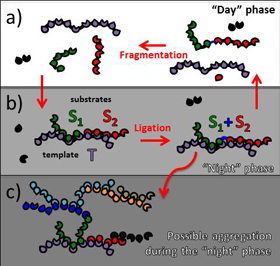
Tkachenko and Maslov have proposed a new model that shows how the earliest self-replicating molecules could have worked. Their model switches between “day” phases, where individual polymers float freely, and “night” phases, where they join together to form longer chains via template-assisted ligation. The phases are driven by cyclic changes in environmental conditions, such as temperature, pH, or salinity, which throw the system out of equilibrium and induce the polymers to either come together or drift apart.
According to their model, during the night cycles, multiple short polymers bond to longer polymer strands, which act as templates. These longer template strands hold the shorter polymers in close enough proximity to each other that they can ligate to form a longer strand — a complementary copy of at least part of the template. Over time, the newly synthesized polymers come to dominate, giving rise to an autocatalytic and self-sustaining system of molecules large enough to potentially encode blueprints for life, the model predicts.
Polymers can also link together without the aid of a template, but the process is somewhat more random — a chain that forms in one generation will not necessarily be carried over into the next. Template-assisted ligation, on the other hand, is a more faithful means of preserving information, as the polymer chains of one generation are used to build the next. Thus, a model based on template-assisted ligation combines the lengthening of polymer chains with their replication, providing a potential mechanism for heritability.
While some previous studies have argued that a mix of the two is necessary for moving a system from monomers to self-replicating polymers, Maslov and Tkachenko’s model demonstrates that it is physically possible for self-replication to emerge with only template-assisted ligation.
“What we have demonstrated for the first time is that even if all you have is template-assisted ligation, you can still bootstrap the system out of primordial soup,” said Maslov.
The idea of template-assisted ligation driving self-replication was originally proposed in the 1980s, but in a qualitative manner. “Now it’s a real model that you can run through a computer,” said Tkachenko. “It’s a solid piece of science to which you can add other features and study memory effects and inheritance.”
Under Tkachenko and Maslov’s model, the move from monomers to polymers is a very sudden one. It’s also hysteretic — that is, it takes a very certain set of conditions to make the initial leap from monomers to self-replicating polymers, but those stringent requirements are not necessary to maintain a system of self-replicating polymers once one has leapt over the first hurdle.
 One limitation of the model that the researchers plan to address in future studies is its assumption that all polymer sequences are equally likely to occur. Transmission of information requires heritable variation in sequence frequencies — certain combinations of bases code for particular proteins, which have different functions. The next step, then, is to consider a scenario in which some sequences become more common than others, allowing the system to transmit meaningful information.
One limitation of the model that the researchers plan to address in future studies is its assumption that all polymer sequences are equally likely to occur. Transmission of information requires heritable variation in sequence frequencies — certain combinations of bases code for particular proteins, which have different functions. The next step, then, is to consider a scenario in which some sequences become more common than others, allowing the system to transmit meaningful information.
Maslov and Tkachenko’s model fits into the currently favored RNA world hypothesis — the belief that life on earth started with autocatalytic RNA molecules that then lead to the more stable but more complex DNA as a mode of inheritance. But because it is so general, it could be used to test any origins of life hypothesis that relies on the emergence of a simple autocatalytic system.
“The model, by design, is very general,” said Maslov. “We’re not trying to address the question of what this primordial soup of monomers is coming from” or the specific molecules involved. Rather, their model shows a physically plausible path from monomer to self-replicating polymer, inching a step closer to understanding the origins of life.
###
BACKGROUNDER: Waiter, there’s an RNA in my Primordial Soup — Tracing the Origins of Life, from Darwin to Today
Nearly every culture on earth has an origins story, a legend explaining its existence. We humans seem to have a deep need for an explanation of how we ended up here, on this small planet spinning through a vast universe. Scientists, too, have long searched for our origins story, trying to discern how, on a molecular scale, the earth shifted from a mess of inorganic molecules to an ordered system of life. The question is impossible to answer for certain — there’s no fossil record, and no eyewitnesses. But that hasn’t stopped scientists from trying.
Over the past 150 years, our shifting understanding of the origins of life has mirrored the emergence and development of the fields of organic chemistry and molecular biology. That is, increased understanding of the role that nucleotides, proteins and genes play in shaping our living world today has also gradually improved our ability to peer into their mysterious past.
When Charles Darwin published his seminal On the Origin of the Species in 1859, he said little about the emergence of life itself, possibly because, at the time, there was no way to test such ideas. His only real remarks on the subject come from a later letter to a friend, in which he suggested a that life emerged out of a “warm little pond” with a rich chemical broth of ions. Nevertheless, Darwin’s influence was far-reaching, and his offhand remark formed the basis of many origins of life scenarios in the following years.
In the early 20th century, the idea was popularized and expanded upon by a Russian biochemist named Alexander Oparin. He proposed that the atmosphere on the early earth was reducing, meaning it had an excess of negative charge. This charge imbalance could catalyze existing a prebiotic soup of organic molecules into the earliest forms of life.
Oparin’s writing eventually inspired Harold Urey, who began to champion Oparin’s proposal. Urey then caught the attention of Stanley Miller, who decided to formally test the idea. Miller took a mixture of what he believed the early earth’s oceans may have contained — a reducing mixture of methane, ammonia, hydrogen, and water — and activated it with an electric spark. The jolt of electricity, acting like a strike of lightening, transformed nearly half of the carbon in the methane into organic compounds. One of the compounds he produced was glycine, the simplest amino acid.
The groundbreaking Miller-Urey experiment showed that inorganic matter could give rise to organic structures. And while the idea of a reducing atmosphere gradually fell out of favor, replaced by an environment rich in carbon dioxide, Oparin’s basic framework of a primordial soup rich with organic molecules stuck around.
The identification of DNA as the hereditary material common to all life, and the discovery that DNA coded for RNA, which coded for proteins, provided fresh insight into the molecular basis for life. But it also forced origins of life researchers to answer a challenging question: how could this complicated molecular machinery have started? DNA is a complex molecule, requiring a coordinated team of enzymes and proteins to replicate itself. Its spontaneous emergence seemed improbable.
In the 1960s, three scientists — Leslie Orgel, Francis Crick and Carl Woese — independently suggested that RNA might be the missing link. Because RNA can self-replicate, it could have acted as both the genetic material and the catalyst for early life on earth. DNA, more stable but more complex, would have emerged later.
Today, it is widely believed (though by no means universally accepted) that at some point in history, an RNA-based world dominated the earth. But how it got there — and whether there was a simpler system before it — is still up for debate. Many argue that RNA is too complicated to have been the first self-replicating system on earth, and that something simpler preceded it.
Graham Cairns-Smith, for instance, has argued since the 1960s that the earliest gene-like structures were not based on nucleic acids, but on imperfect crystals that emerged from clay. The defects in the crystals, he believed, stored information that could be replicated and passed from one crystal to another. His idea, while intriguing, is not widely accepted today.
Others, taken more seriously, suspect that RNA may have emerged in concert with peptides — an RNA-peptide world, in which the two worked together to build up complexity. Biochemical studies are also providing insight into simpler nucleic acid analogs that could have preceded the familiar bases that make up RNA today. It’s also possible that the earliest self-replicating systems on earth have left no trace of themselves in our current biochemical systems. We may never know, and yet, the challenge of the search seems to be part of its appeal.
Recent research by Tkachenko and Maslov, published July 28, 2015 in The Journal of Chemical Physics, suggests that self-replicating molecules such as RNA may have arisen through a process called template-assisted ligation. That is, under certain environmental conditions, small polymers could be driven to bond to longer complementary polymer template strands, holding the short strands in close enough proximity to each other that they could fuse into longer strands. Through cyclic changes in environmental conditions that induce complementary strands to come together and then fall apart repeatedly, a self-sustaining collection of hybridized, self-replicating polymers able to encode the blueprints for life could emerge.
###
For More Information:
Jason Socrates Bardi
jbardi@aip.org
240-535-4954
@jasonbardi
Article Title
Spontaneous emergence of autocatalytic information-coding polymers
Authors
Alexei Tkachenko and Sergei Maslov
Author Affiliations
Brookhaven National Laboratory and University of Illinois at Urbana-Champaign
